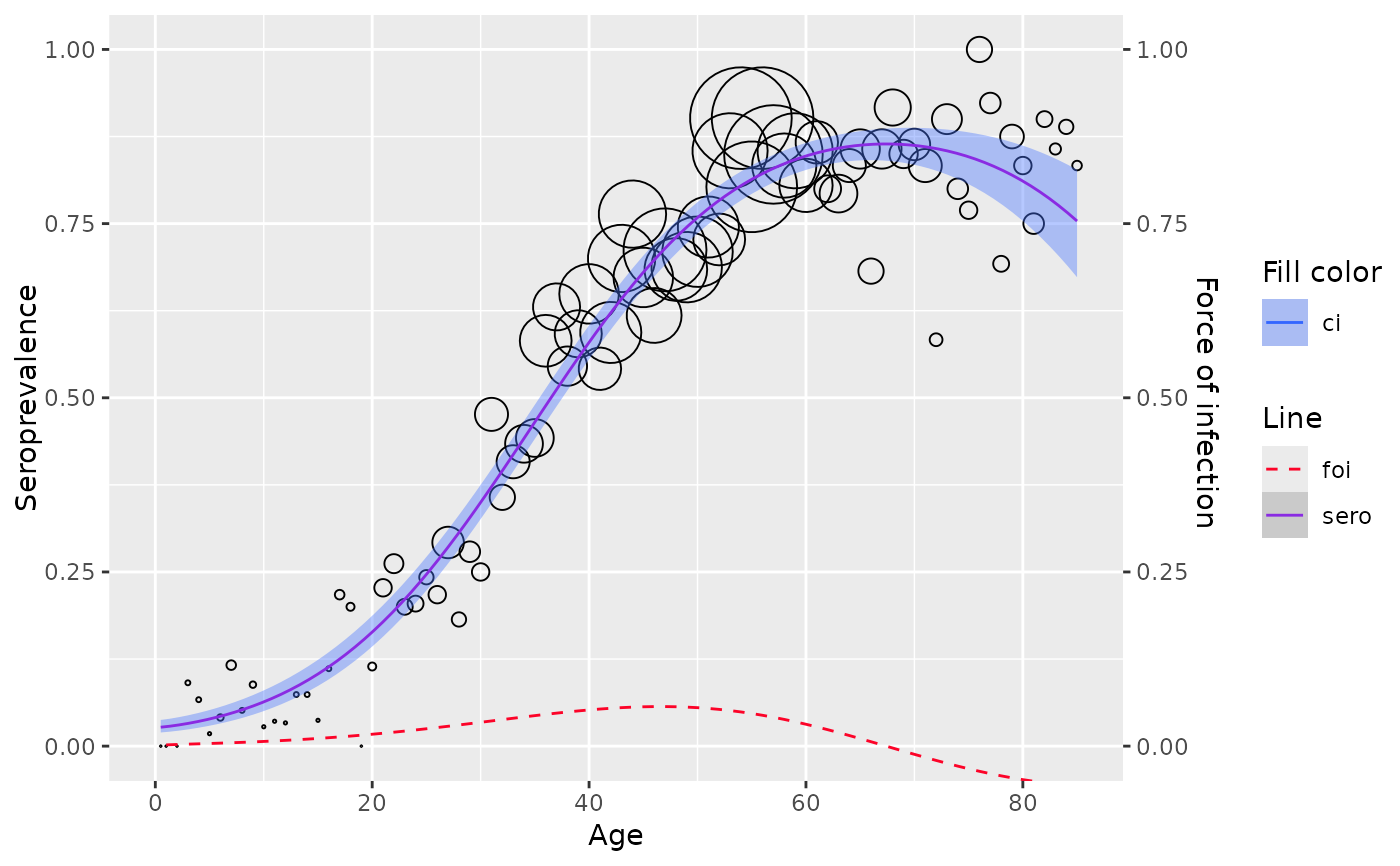
Fractional polynomial model is a generalization of polynomial models where the power of the terms can be fractions, allowing more flexibility and better fit for data where asymptotic behavior is expected.
Usage
fp_model(
data,
p,
monotonic = FALSE,
link = "logit",
age_col = "age",
pos_col = "pos",
tot_col = "tot",
status_col = "status"
)Arguments
- data
the input data frame, must either have `age`, `pos`, `tot` columns (for aggregated data) OR `age`, `status` for (linelisting data)
- p
is either:
a numeric vector specifying the powers to apply to the predictors
a named list with two elements,
"p_range"and"degree"."p_range"is a sequence of powers and"degree"is the maximum degree. In which case the package will search for the best degree and power combinations
- monotonic
whether the returned model should be monotonic (if a search is specified)
- link
the link function for model. Defaulted to "logit".
- age_col
name of the `age` column (default age_col="age").
- pos_col
name of the `pos` column (default pos_col="pos").
- tot_col
name of the `tot` column (default tot_col="tot").
- status_col
name of the `status` column (default status_col="status").
Value
a list of class fp_model with 5 items
- datatype
type of data used for fitting model (aggregated or linelisting)
- df
the dataframe used for fitting the model
- info
a fitted glm model
- sp
seroprevalence
- foi
force of infection
Details
Instead of a polynomial, the linear predictor is now defined as $$ \eta_m(a, \beta, p_1, p_2, ...,p_m) = \Sigma^m_{i=0} \beta_i H_i(a) $$ Where \(m\) is an integer, \(p_1 \le p_2 \le... \le p_m\) is a sequence of powers, and \(H_i(a)\) is a transformation given by
$$ H_i = \begin{cases} a^{p_i} & \text{ if } p_i \neq p_{i-1}, \\ H_{i-1}(a) \times log(a) & \text{ if } p_i = p_{i-1}, \end{cases} $$
Refers to section 6.2. of the the book by Hens et al. (2012) for further details.
References
Hens, Niel, Ziv Shkedy, Marc Aerts, Christel Faes, Pierre Van Damme, and Philippe Beutels. 2012. Modeling Infectious Disease Parameters Based on Serological and Social Contact Data: A Modern Statistical Perspective. tatistics for Biology and Health. Springer New York. doi:10.1007/978-1-4614-4072-7 .