Fit age-stratified seroprevalence to parametric hierarchical Bayesian models. Supported models including Farrington model (2 and 3 parameters variants) and Log Logistic model
Usage
hierarchical_bayesian_model(
data,
age_col = "age",
pos_col = "pos",
tot_col = "tot",
status_col = "status",
type = "far3",
chains = 1,
warmup = 1500,
iter = 5000
)Arguments
- data
the input data frame, must either have columns for `age`, `pos`, `tot` (for aggregated data) OR `age`, `status` (for linelisting data)
- age_col
name of the `age` column (default age_col="age").
- pos_col
name of the `pos` column (default pos_col="pos").
- tot_col
name of the `tot` column (default tot_col="tot").
- status_col
name of the `status` column (default status_col="status").
- type
type of model ("far2", "far3" or "log_logistic")
- chains
number of Markov chains
- warmup
number of warmup runs
- iter
number of iterations
Value
a list of class hierarchical_bayesian_model with 6 items
- datatype
type of datatype used for model fitting (aggregated or linelisting)
- df
the dataframe used for fitting the model
- type
type of bayesian model far2, far3 or log_logistic
- info
parameters for the fitted model
- sp
seroprevalence
- foi
force of infection
- sp_func
function to compute seroprevalence given age and model parameters
- foi
function to compute force of infection given age and model parameters
Details
Consider a model for prevalence that has a parametric form \(\pi(a_i, \alpha)\) where \(\alpha\) is a parameter vector
Under a Bayesian framework, we can constraint the parameter space of the prior distribution \(P(\alpha)\) to achieve monotonicity of the posterior distribution \(P(\pi_1, \pi_2, ..., \pi_m|y,n)\)
Where:
- \(n = (n_1, n_2, ..., n_m)\) and \(n_i\) is the sample size at age \(a_i\)
- \(y = (y_1, y_2, ..., y_m)\) and \(y_i\) is the number of infected individual from the \(n_i\) sampled subjects
For Farrington model with 3 parameters, prevalence is formulated as follow
$$ \pi (a) = 1 - exp\{ \frac{\alpha_1}{\alpha_2}ae^{-\alpha_2 a} + \frac{1}{\alpha_2}(\frac{\alpha_1}{\alpha_2} - \alpha_3)(e^{-\alpha_2 a} - 1) -\alpha_3 a \} $$
The likelihood model is defined as \(y_i \sim Bin(n_i, \pi_i), \text{ for } i = 1,2,3,...m\)
The constraint on the parameter space can be incorporated by assuming truncated normal distribution for the components of \(\alpha\), \(\alpha = (\alpha_1, \alpha_2, \alpha_3)\) in \(\pi_i = \pi(a_i,\alpha)\)
The flat hyperpriors are defined as \(\mu_j \sim \mathcal{N}(0, 10000)\) and \(\tau^{-2}_j \sim \Gamma(100,100)\)
For Farrington model with 2 parameters, it is equivalent to the previous model with \(\alpha_3 = 0\)
For Log logistic model, seroprevalence is instead defined as
$$\pi(a) = \frac{\beta a^\alpha}{1 + \beta a^\alpha}, \text{ } \alpha, \beta > 0$$
The likelihood is similarly defined as \(y_i \sim Bin(n_i, \pi_i))\)
The prior model of \(\alpha_1\) is specified as \(\alpha_1 \sim \text{truncated } \mathcal{N}(\mu_1, \tau_1)\) with flat hyperpriors as in Farrington model
\(\beta\) is constrained to be positive by specifying \(\alpha_2 \sim \mathcal{N}(\mu_2, \tau_2)\)
Refer to section Chapter 10.3 of the the book by Hens et al. (2012) for further details.
References
Hens, Niel, Ziv Shkedy, Marc Aerts, Christel Faes, Pierre Van Damme, and Philippe Beutels. 2012. Modeling Infectious Disease Parameters Based on Serological and Social Contact Data: A Modern Statistical Perspective. tatistics for Biology and Health. Springer New York. doi:10.1007/978-1-4614-4072-7 .
Examples
# \donttest{
df <- mumps_uk_1986_1987
model <- hierarchical_bayesian_model(df, type="far3")
#>
#> SAMPLING FOR MODEL 'fra_3' NOW (CHAIN 1).
#> Chain 1: Rejecting initial value:
#> Chain 1: Log probability evaluates to log(0), i.e. negative infinity.
#> Chain 1: Stan can't start sampling from this initial value.
#> Chain 1: Rejecting initial value:
#> Chain 1: Log probability evaluates to log(0), i.e. negative infinity.
#> Chain 1: Stan can't start sampling from this initial value.
#> Chain 1: Rejecting initial value:
#> Chain 1: Log probability evaluates to log(0), i.e. negative infinity.
#> Chain 1: Stan can't start sampling from this initial value.
#> Chain 1: Rejecting initial value:
#> Chain 1: Log probability evaluates to log(0), i.e. negative infinity.
#> Chain 1: Stan can't start sampling from this initial value.
#> Chain 1:
#> Chain 1: Gradient evaluation took 0.00013 seconds
#> Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 1.3 seconds.
#> Chain 1: Adjust your expectations accordingly!
#> Chain 1:
#> Chain 1:
#> Chain 1: Iteration: 1 / 5000 [ 0%] (Warmup)
#> Chain 1: Iteration: 500 / 5000 [ 10%] (Warmup)
#> Chain 1: Iteration: 1000 / 5000 [ 20%] (Warmup)
#> Chain 1: Iteration: 1500 / 5000 [ 30%] (Warmup)
#> Chain 1: Iteration: 1501 / 5000 [ 30%] (Sampling)
#> Chain 1: Iteration: 2000 / 5000 [ 40%] (Sampling)
#> Chain 1: Iteration: 2500 / 5000 [ 50%] (Sampling)
#> Chain 1: Iteration: 3000 / 5000 [ 60%] (Sampling)
#> Chain 1: Iteration: 3500 / 5000 [ 70%] (Sampling)
#> Chain 1: Iteration: 4000 / 5000 [ 80%] (Sampling)
#> Chain 1: Iteration: 4500 / 5000 [ 90%] (Sampling)
#> Chain 1: Iteration: 5000 / 5000 [100%] (Sampling)
#> Chain 1:
#> Chain 1: Elapsed Time: 15.235 seconds (Warm-up)
#> Chain 1: 139.842 seconds (Sampling)
#> Chain 1: 155.077 seconds (Total)
#> Chain 1:
#> Warning: There were 324 divergent transitions after warmup. See
#> https://mc-stan.org/misc/warnings.html#divergent-transitions-after-warmup
#> to find out why this is a problem and how to eliminate them.
#> Warning: There were 143 transitions after warmup that exceeded the maximum treedepth. Increase max_treedepth above 10. See
#> https://mc-stan.org/misc/warnings.html#maximum-treedepth-exceeded
#> Warning: Examine the pairs() plot to diagnose sampling problems
#> Warning: Bulk Effective Samples Size (ESS) is too low, indicating posterior means and medians may be unreliable.
#> Running the chains for more iterations may help. See
#> https://mc-stan.org/misc/warnings.html#bulk-ess
#> Warning: Tail Effective Samples Size (ESS) is too low, indicating posterior variances and tail quantiles may be unreliable.
#> Running the chains for more iterations may help. See
#> https://mc-stan.org/misc/warnings.html#tail-ess
model$info
#> mean se_mean sd 2.5%
#> alpha1 1.394994e-01 2.208144e-04 5.686602e-03 1.290184e-01
#> alpha2 1.984613e-01 3.072933e-04 7.829686e-03 1.844645e-01
#> alpha3 8.289398e-03 2.217734e-04 6.839120e-03 3.133171e-04
#> tau_alpha1 6.381566e+00 1.529942e+00 1.636028e+01 6.597716e-06
#> tau_alpha2 1.371545e+01 8.454139e+00 4.030987e+01 5.871040e-06
#> tau_alpha3 5.780803e+00 1.085321e+00 1.356397e+01 7.111779e-06
#> mu_alpha1 8.127640e-01 1.294292e+00 2.957688e+01 -7.133897e+01
#> mu_alpha2 3.575058e+00 1.799463e+00 3.726214e+01 -7.253911e+01
#> mu_alpha3 3.883757e+00 1.307763e+00 3.405219e+01 -6.454653e+01
#> sigma_alpha1 6.352800e+01 1.682171e+01 7.004084e+02 1.213045e-01
#> sigma_alpha2 7.855927e+01 1.684436e+01 6.470535e+02 7.736001e-02
#> sigma_alpha3 1.873854e+02 1.323037e+02 3.877173e+03 1.431469e-01
#> lp__ -2.532504e+03 3.710684e-01 4.655392e+00 -2.542452e+03
#> 25% 50% 75% 97.5% n_eff
#> alpha1 1.357106e-01 1.390419e-01 1.430492e-01 1.511064e-01 663.20993
#> alpha2 1.931963e-01 1.983732e-01 2.027147e-01 2.165249e-01 649.20593
#> alpha3 2.841082e-03 6.688870e-03 1.163816e-02 2.583495e-02 951.00250
#> tau_alpha1 1.733407e-03 9.495067e-02 2.687740e+00 6.795883e+01 114.34889
#> tau_alpha2 1.342653e-03 6.464717e-02 2.511530e+00 1.670964e+02 22.73443
#> tau_alpha3 1.111049e-03 5.337217e-02 2.822276e+00 4.880180e+01 156.19150
#> mu_alpha1 -1.276472e+00 1.729980e-01 2.532245e+00 5.703259e+01 522.20403
#> mu_alpha2 -1.528020e+00 2.032933e-01 2.920524e+00 1.045006e+02 428.79515
#> mu_alpha3 -1.524451e+00 3.939297e-02 3.640311e+00 9.311625e+01 678.00364
#> sigma_alpha1 6.099697e-01 3.245271e+00 2.401891e+01 3.893194e+02 1733.65511
#> sigma_alpha2 6.310022e-01 3.933036e+00 2.729092e+01 4.127492e+02 1475.60823
#> sigma_alpha3 5.952566e-01 4.328557e+00 3.000085e+01 3.749841e+02 858.78884
#> lp__ -2.535634e+03 -2.532144e+03 -2.528948e+03 -2.524553e+03 157.39993
#> Rhat
#> alpha1 1.0002098
#> alpha2 0.9998023
#> alpha3 0.9997596
#> tau_alpha1 1.0003182
#> tau_alpha2 1.0444726
#> tau_alpha3 1.0101265
#> mu_alpha1 1.0073358
#> mu_alpha2 1.0048162
#> mu_alpha3 1.0031755
#> sigma_alpha1 0.9997210
#> sigma_alpha2 1.0000959
#> sigma_alpha3 1.0009114
#> lp__ 0.9998177
plot(model)
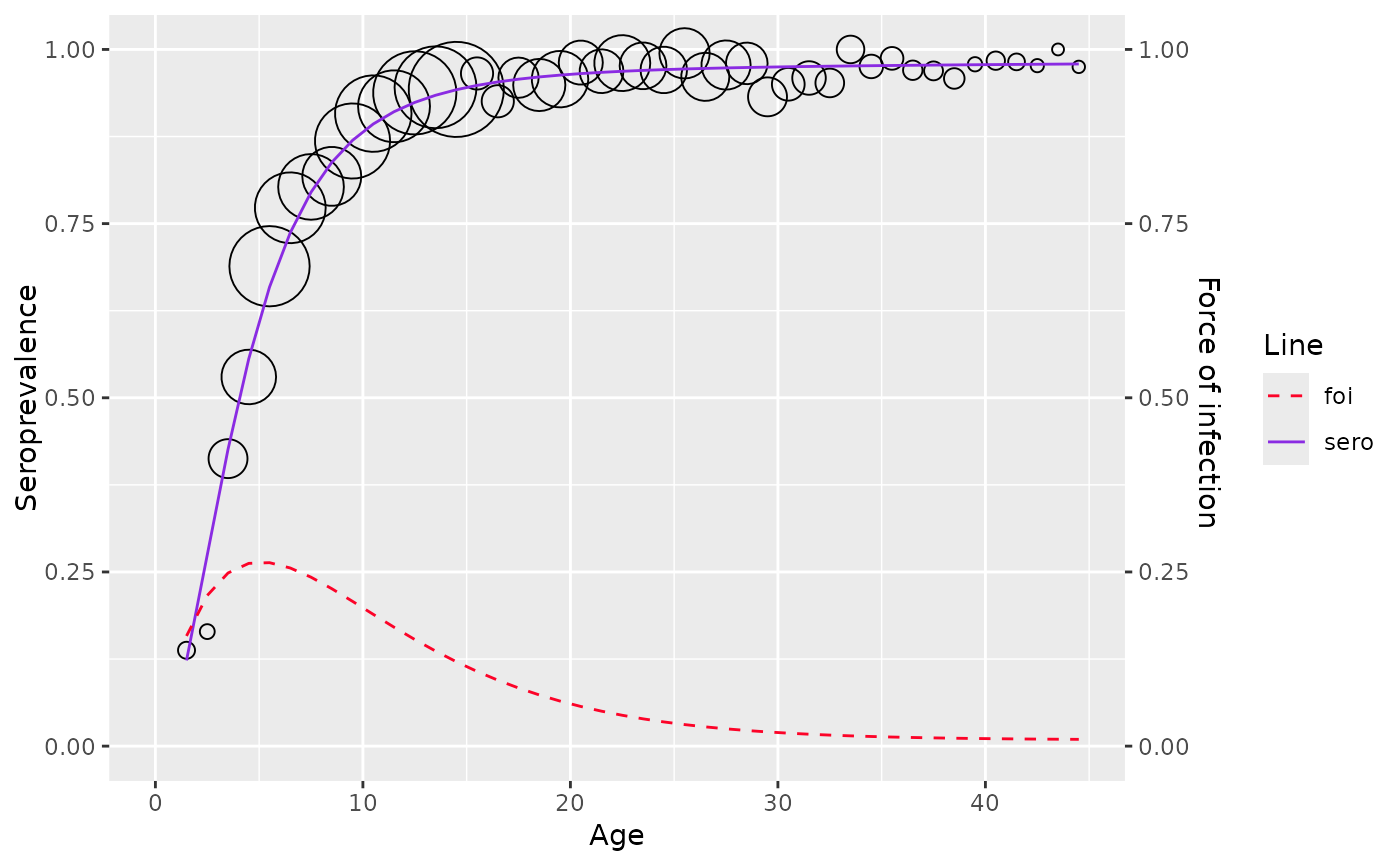 # }
# }