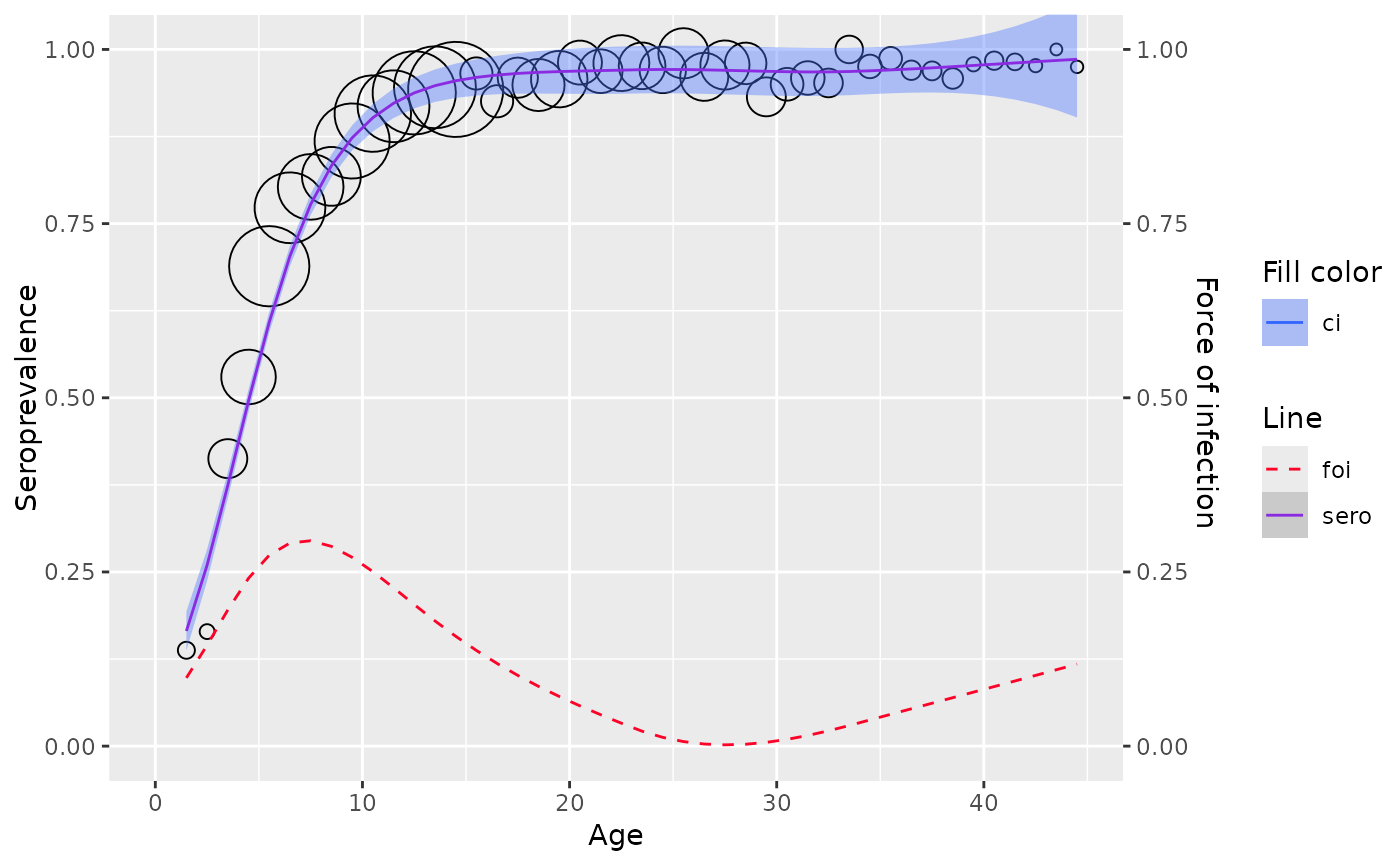
Fit the age-specific seroprevalence to a local polynomial model, where the linear predictor is approximated locally at one particular age.
Usage
lp_model(
data,
kern = "tcub",
nn = 0,
h = 0,
deg = 2,
age_col = "age",
pos_col = "pos",
tot_col = "tot",
status_col = "status"
)Arguments
- data
the input data frame, must either have columns for `age`, `pos`, `tot` (for aggregated data) OR `age`, `status` (for linelisting data)
- kern
Weight function, default = "tcub". Other choices are "rect", "trwt", "tria", "epan", "bisq" and "gauss". Choices may be restricted when derivatives are required; e.g. for confidence bands and some bandwidth selectors.
- nn
Nearest neighbor component of the smoothing parameter. Default value is 0.7, unless either h is provided, in which case the default is 0.
- h
The constant bandwidth of the smoothing parameter. Default: 0.
- deg
Degree of polynomial to use. Default: 2.
- age_col
name of the `age` column (default age_col="age").
- pos_col
name of the `pos` column (default pos_col="pos").
- tot_col
name of the `tot` column (default tot_col="tot").
- status_col
name of the `status` column (default status_col="status").
Value
a list of class lp_model with 6 items
- datatype
type of datatype used for model fitting (aggregated or linelisting)
- df
the dataframe used for fitting the model
- pi
fitted locfit object for pi
- eta
fitted locfit object for eta
- sp
seroprevalence
- foi
force of infection
Details
Consider a linear predictor \(\eta(a)\) approximated locally at one particular value \(a_0\).
For a general degree \(p\), the linear predictor for a neighbor of \(a_0\), labeled \(a_i\) is equivalent to the Taylor approximation $$ \eta(a_i) = \eta(a_0) + \eta^{(1)}(a_0)(a_i - a_0) + \frac{\eta^{(2)}(a_0)}{2}(a_i - a_0)^2 + ... + \frac{\eta^{(p)}(a_0)}{p!}(a_i - a_0)^p $$
\(\eta(a_i)\) can be estimated by maximizing $$ \Sigma_{i=1}^{N} \ell_i \{Y_i, g^{-1} (\beta_0 + \beta_1(a_i-a_0)+ \beta_2(a_i-a_0)^2 ... + \beta_p(a_i-a_0)^p) \} K_h(a_i - a_0) $$
The estimator for the \(k\)-th derivative of \(\eta(a_0)\), for \(k = 0,1,…,p\) (degree of local polynomial) is thus: $$ \hat{\eta}^{(k)}(a_0) = k!\hat{\beta}_k(a_0) $$
The estimator for the prevalence at age \(a_0\) is then given by $$ \hat{\pi}(a_0) = g^{-1}\{ \hat{\beta}_0(a_0) \} $$ Where \(g\) is the link function
The estimator for the force of infection at age \(a_0\) by assuming \(p \ge 1\) is as followed $$ \hat{\lambda}(a_0) = \hat{\beta}_1(a_0) \delta \{ \hat{\beta}_0 (a_0) \} $$ Where \(\delta \{ \hat{\beta}_0(a_0) \} = \frac{dg^{-1} \{ \hat{\beta}_0(a_0) \} } {d\hat{\beta}_0(a_0)}\)
Refer to section 7.1 and 7.2. of the the book by Hens et al. (2012) for further details.
References
Hens, Niel, Ziv Shkedy, Marc Aerts, Christel Faes, Pierre Van Damme, and Philippe Beutels. 2012. Modeling Infectious Disease Parameters Based on Serological and Social Contact Data: A Modern Statistical Perspective. tatistics for Biology and Health. Springer New York. doi:10.1007/978-1-4614-4072-7 .
Examples
df <- mumps_uk_1986_1987
model <- lp_model(
df,
nn=0.7, kern="tcub"
)
plot(model)