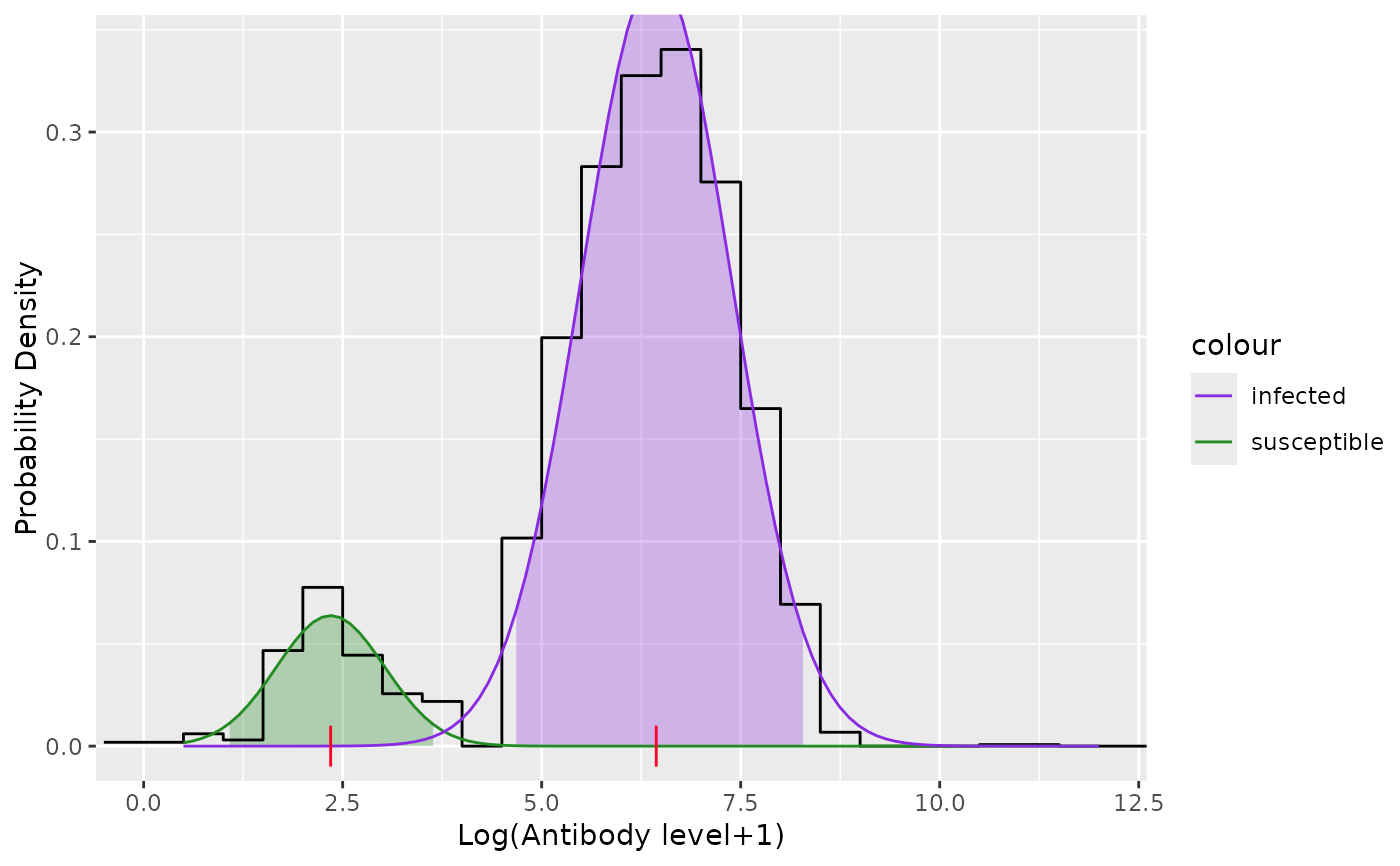
Fit the antibody level data to a 2-component Gaussian mixture model
Arguments
- antibody_level
vector of the corresponding raw antibody level
- breaks
number of intervals which the antibody_level are grouped into
- pi
proportion of susceptible, infected
- mu
a vector of means of component distributions (vector of 2 numbers in ascending order)
- sigma
a vector of standard deviations of component distributions (vector of 2 number)
Value
a list of class mixture_model with the following items
- df
the dataframe used for fitting the model
- info
list of 3 items parameters, distribution and constraints for the fitted model
- susceptible
fitted distribution for susceptible
- infected
fitted distribution for infected
Details
Antibody level (denoted \(Z\)) is modeled using a 2-component Gaussian mixture model. Each component \(Z_j\) (\(j \in \{I, S\}\)) represents the antibody level of the latent Infected and Susceptible sub-populations, following density \(f_j(z_j|\theta_j)\)
Let \(\pi_{\text{TRUE}}(a)\) denotes the age-dependent mixing probability (i.e., the true prevalence), the density of the mixture is formulated as
$$f(z|z_I, z_S,a) = (1-\pi_{\text{TRUE}}(a))f_S(z_S|\theta_S)+\pi_{\text{TRUE}}(a)f_I(z_I|\theta_I)$$
The mean \(E(Z|a)\) thus equals $$\mu(a) = (1-\pi_{\text{TRUE}}(a))\mu_S+\pi_{\text{TRUE}}(a)\mu_I$$
From which true prevalence can be computed as $$\pi_{\text{TRUE}}(a) = \frac{\mu(a) - \mu_S}{\mu_I - \mu_S}$$
And FOI can then be inferred as $$\lambda_{TRUE} = \frac{\mu'(a)}{\mu_I - \mu(a)}$$
Function [serosv::mixture_model()] fits antibody level data to \(f_S(z_S|\theta_S)\) and \(f_I(z_I|\theta_I)\)
Function [serosv::estimate_mixture()] will then estimate age-specific antibody level \(\mu(a)\) and infer the estimation for \(\pi_{\text{TRUE}}(a)\) and \(\lambda_{TRUE}\)
Refer to section 11.3. of the the book by Hens et al. (2012) for further details.
References
Hens, Niel, Ziv Shkedy, Marc Aerts, Christel Faes, Pierre Van Damme, and Philippe Beutels. 2012. Modeling Infectious Disease Parameters Based on Serological and Social Contact Data: A Modern Statistical Perspective. tatistics for Biology and Health. Springer New York. doi:10.1007/978-1-4614-4072-7 .
Examples
df <- vzv_be_2001_2003[vzv_be_2001_2003$age < 40.5,]
data <- df$VZVmIUml[order(df$age)]
model <- mixture_model(antibody_level = data)
model$info
#>
#> Parameters:
#> pi mu sigma
#> 1 0.1088 2.349 0.6804
#> 2 0.8912 6.439 0.9437
#>
#> Distribution:
#> [1] "norm"
#>
#> Constraints:
#> conpi conmu consigma
#> "NONE" "NONE" "NONE"
#>
plot(model)