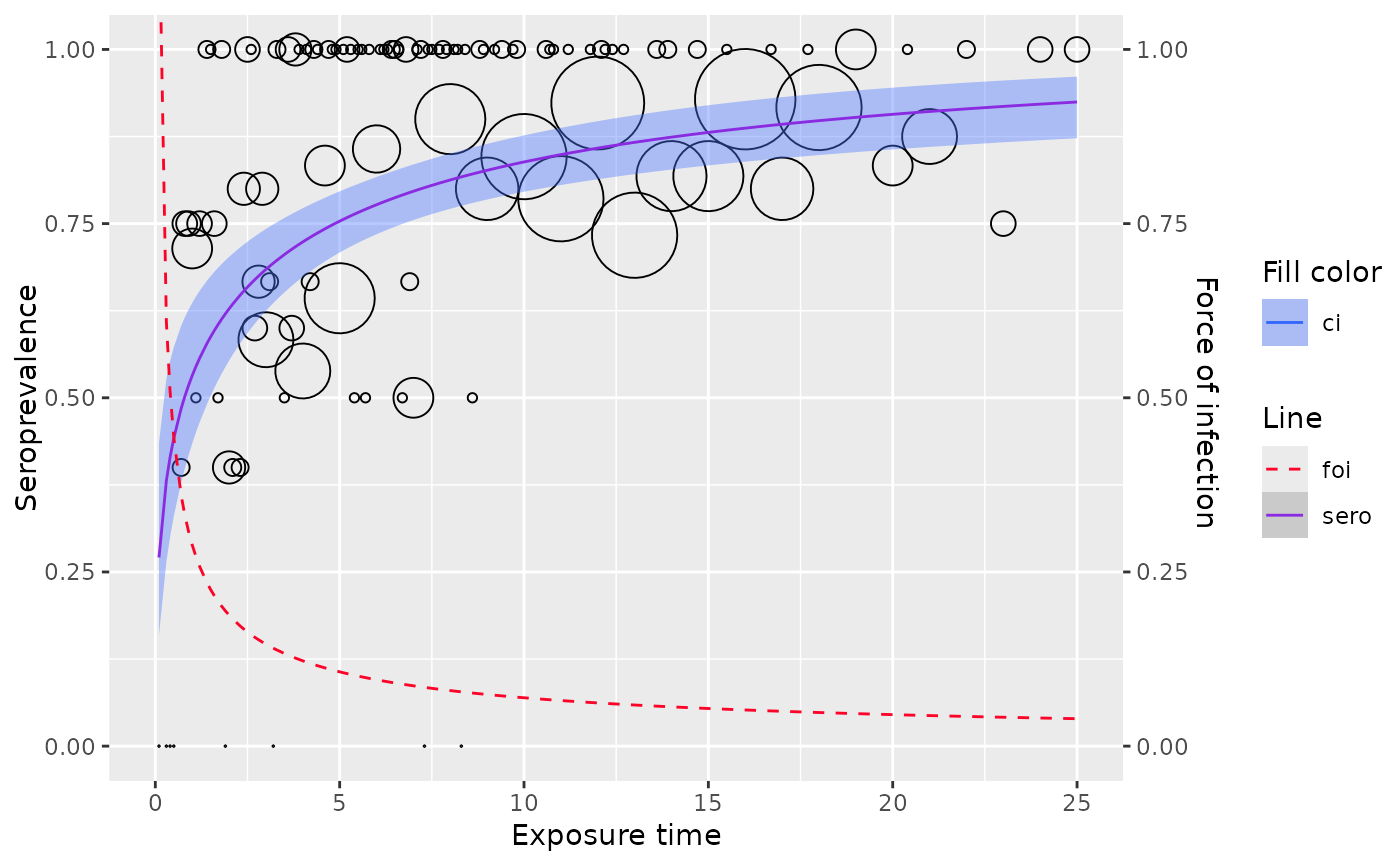
Model seroprevalence as a function of duration since vaccination using the Weibull model, where the force of infection is assumed to vary monotonically with duration.
Arguments
- data
the input data frame, must either have columns for `t`, `pos`, `tot` (for aggregated data) OR `t`, `status` (for linelisting data)
- t_lab
name of the `t` column (default t_lab="t").
- pos_col
name of the `pos` column (default pos_col="pos").
- tot_col
name of the `tot` column (default tot_col="tot").
- status_col
name of the `status` column (default status_col="status").
Value
list of class weibull_model with the following items
- datatype
type of datatype used for model fitting (aggregated or linelisting)
- df
the dataframe used for fitting the model
- info
fitted "glm" object
- sp
seroprevalence
- foi
force of infection
Details
For a Weibull model, the prevalence is given by $$ \pi (d) = 1 - e^{ - \beta_0 d ^ {\beta_1}} $$ Where \(d\) is exposure time (difference between age of vaccination and age at test)
Which implies the force of infection to be the monotonic function $$ \lambda(d) = \beta_0 \beta_1 d^{\beta_1 - 1} $$
Refer to section 6.1.2. of the the book by Hens et al. (2012) for further details.
References
Hens, Niel, Ziv Shkedy, Marc Aerts, Christel Faes, Pierre Van Damme, and Philippe Beutels. 2012. Modeling Infectious Disease Parameters Based on Serological and Social Contact Data: A Modern Statistical Perspective. tatistics for Biology and Health. Springer New York. doi:10.1007/978-1-4614-4072-7 .